COEVOL Multi-Scale Coevolution
Living systems are highly integrated, with a multitude of levels of organization, from molecular and intra-cellular scales to ecosystems. Complex organisms are themselves consortia of macro- and micro-organisms, which work together with their host to build the individual. Yet, each of these organisms can function and evolve in the short term according to its own logic, possibly in conflict with other higher or lower levels, or with other time scales. The once common idea among evolutionists that natural selection results in organisms perfectly adapted to their environment is now severely undermined. Not only because, as the Red Queen explains to Alice, one has to run relentlessly to keep its place in a changing environment, or because past evolutionary history and chance constrain the possibilities of present adaptation, but also because different levels of selection have interests that are generally difficult to reconcile.
Multi-scale coevolution resets classical questions in evolutionary biology
One example, of particular interest is the question of the source of heritable variations. The phenotype of organisms in a population is influenced not only by variations in their nuclear and mitochondrial genomes, the dynamics of which is the object of population genetics, but also more and more patently by the consortium of microbes and genetic elements that constitute its microbiome and virome. The hologenome designates this complex assembly of genetic materials, which obey different rules of transmission and different evolutionary strategies. The ability of symbionts to manipulate host phenotypes or to interfere with each other influences the evolutionary dynamics of all players in ways that are yet poorly understood. In addition, new questions arise, such as the importance of co-adaptation in these systems and their consequences in maintaining cohesive biological systems.
- Symbiosis: a response to and a source of divergent selection
Using a variety of approaches combining experimental evolution, genomic, functional, phenotypic and behavioral data, we aim to test whether symbiosis facilitates diversification and to characterize the underlying microevolutionary processes.
- Ecological networks of horizontal gene transfer
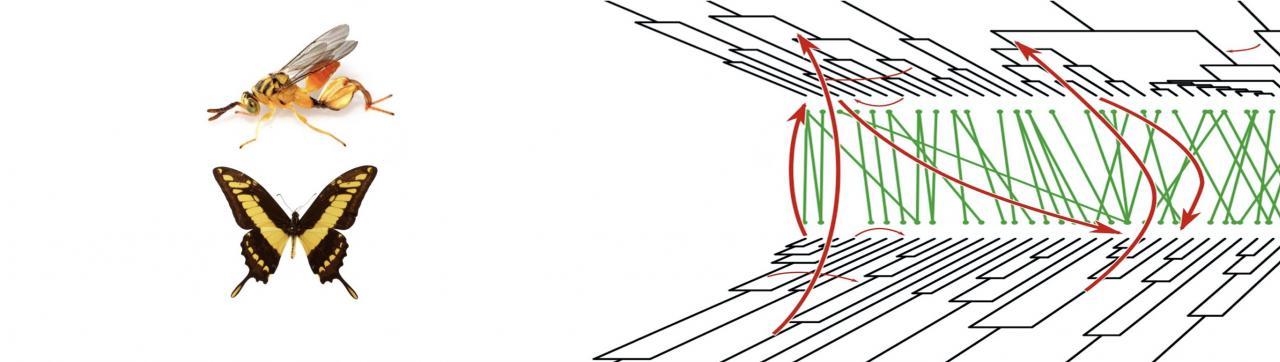
We develop original methods to detect gene transfer and we investigate the factors that influence the routes of gene transfers among microbes but also among insects.
- The interplay between symbiosis, infection and immunity and its evolutionary consequences
We try to understand the intimate interaction of hosts with pathogens, symbionts and transposable elements and how it affects the extended phenotype of the host.
- Transgenerational inheritance and environment changes
We try to decipher the molecular mechanisms that underlie rapid adaptation to environment and to test for transgenerational inheritance of fitness traits.
- Intragenomic conflicts and demography
We are developing models to test whether changes in the demography of the host affect the dynamics of transposable elements.
- The determinism of phenotypic convergence
We study the genomic basis of convergent phenotypic evolution in particular in the case of animals and plants adaptation to increasing temperature and decreasing water.
- Reconciling the tree of life
We develop phylogenetic methods for “reconciling” gene/species or host/symbiont histories and use these methods to explore the bulk of extinct or undescribed species and the history of association of symbiotic microbes with their hosts.
Integrating methods
The methods we use to tackle the questions raised by multi-scale co-evolution extend from theory, modelling and simulation to big data analysis, lab (notably on insects), and to a lesser extent, field activities.
Implication of research, responsibility of researchers and citizen sciences
From our research (some of which have immediate consequences in health, agriculture and ecology) and our concerns about the responsibility of scientists in society, we are committed to promote an “implicative” research. The implicative position means that we try to work on the link between science and society, not only through a one-way communication, applying or explaining our science, but also favoring early discussions on research projects, that may influence our research directions.
Publications
Display of 511 to 540 publications on 709 in total
Computation of Perfect DCJ Rearrangement Scenarios with Linear and Circular Chromosomes
Journal of Computational Biology . 16 ( 10 ) : 1287-1309
Journal article
see the publicationIdentification of expressed transposable element insertions in the sequenced genome of Drosophila melanogaster
Gene . 439 : 55-62
Journal article
see the publicationMonoallelic expression and tissue specificity are associated with high crossover rates
Trends in Genetics . 25(12) : 519-522
Journal article
see the publicationMaintenance of adaptive differentiation by Wolbachia induced bidirectional cytoplasmic incompatibility: the importance of sib-mating and genetic systems.
BMC Evolutionary Biology . 9 : 185
Journal article
see the publicationHybrid sterility and inviability in the parasitic fungal species complex Microbotryum
Journal of Evolutionary Biology . 22 ( 4 ) : 683--698
Journal article
see the publicationThe relationship between DNA replication and human genome organization.
Molecular Biology and Evolution . 26 ( 4 ) : 729-741
Journal article
see the publicationA Virus-Shaping Reproductive Strategy in a Drosophila Parasitoid
Advances in Parasitology . 70 : 334-359
Journal article
see the publicationMolecular Detection Penetrance and Transmission of an Inherited Virus Responsible for Behavioral Manipulation of an Insect Parasitoid
Applied and Environmental Microbiology . 75 ( 3 ) : 703-710
DOI: 10.1128/AEM.01778-08
Journal article
see the publicationWolbachia interferes with ferritin expression and iron metabolism in insects.
PLoS Pathogens . 5 ( 10 ) : e1000630
Journal article
see the publicationImmunity and symbiosis.
Molecular Microbiology . 73 ( 5 ) : 751-9
Journal article
see the publicationInteractions between vertically transmitted symbionts: cooperation or conflict ?
Trends in Microbiology . 17 : 95-99
Journal article
see the publicationEvolution des interactions symbiotiques chez les insectes : A quel niveau de sélection se vouer ?
incollection . -- : 33-35
Journal article
see the publicationEviter les tricheurs: mode de transmission et stabilité des symbioses
Biofutur . 299 : 33-35
Journal article
see the publicationDrosophila-parasitoid communities as model systems for host-Wolbachia interactions.
Advances in Parasitology . 70 : 299-331
Journal article
see the publicationInfecciones múltiples regulación y evolución de poblaciones simbióticas
incollection . -- : 197-226
Journal article
see the publicationDrosophila–Parasitoid Communities as Model Systems for Host–Wolbachia Interactions
incollection . 70 : 229-331
Journal article
see the publicationPhylogenetic Relationships Deduced from Whole Genome Comparisons
Encyclopedia of Life Sciences . : 1-8
Interactions between Coexisting Intracellular Genomes: Mitochondrial Density and Wolbachia Infection
Applied and Environmental Microbiology . 75(7) : 1916-1921
Journal article
see the publicationGenomic environment influences the dynamics of the tirant LTR retrotransposon in Drosophila.
FASEB Journal . 23 ( 5 ) : 1482-9
DOI: 10.1096/fj.08-123513
Journal article
see the publicationEvolutionary Origin and Functions of Retrogene Introns
Molecular Biology and Evolution . 26(9) : 2147-2156
Journal article
see the publicationHow sympatric is speciation in the Howea palms of Lord Howe Island ?
Molecular Ecology . 18 : 3629-3638
Journal article
see the publicationPhylogenetic determinants of potential host shifts in fungal pathogens
Journal of Evolutionary Biology . 22 ( 12 ) : 2532--2541
Journal article
see the publicationPrediction of Contiguous Regions in the Amniote Ancestral Genome
ISBRA 2009, 5th International Symposium on Bioinformatics Research and Applications, . 5542
Conference paper
see the publicationMultichromosomal median and halving problems under different genomic distances
BMC Bioinformatics . 10 ( 1 ) : 120
Journal article
see the publicationThe evolutionary dynamics of the Helena retrotransposon revealed by sequenced Drosophila genomes.
BMC Evolutionary Biology . 9 : 174
Journal article
see the publicationDatabases of Homologous Gene Families for Comparative Genomics
BMC Bioinformatics . 10 (Suppl.6) :S3 : 13
Journal article
see the publication[Prophylaxis of thromboembolic disease in obstetrics after the French guidelines publication]
Gynécologie Obstétrique & Fertilité . 36 ( 1 ) : 23-34
Journal article
see the publicationDo vertically transmitted symbionts co-existing in a single host compete or cooperate? A modelling approach
Journal of Evolutionary Biology . 21 : 145-161
Journal article
see the publicationInherited intracellular ecosystem: symbiotic bacteria share bacteriocytes in whiteflies
FASEB Journal . 22 : 1-9
Journal article
see the publication